
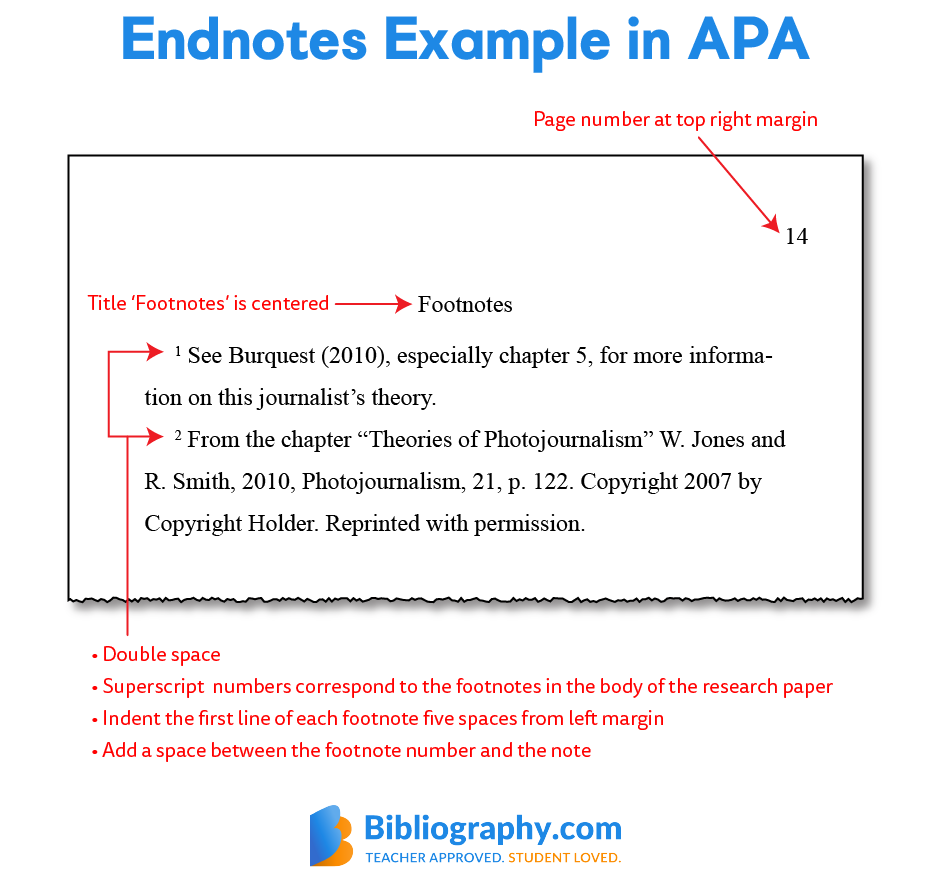
Cancer cells commonly rely on extracellular proteins to provide amino acids. identified LYSET as being selectively essential when cells feed on extracellular proteins. In cells lacking LYSET, the trafficking of enzymes to the lysosomes was severely disrupted, resulting in the accumulation of undigested material in the lysosome. identified a small protein named LYSET that is critical for proper lysosomal function. Certain viruses, including severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2), hijack lysosomes to gain entry into the cell and start their destructive infection cycle. Lysosomes are major degradative compartments within the cell, and their dysfunction results in both rare and common disorders.

This approach allowed for the design of a range of experimentally validated cyclic oligomers and paves the way for the design of increasingly complex assemblies. Although the designs were generated to achieve stable expression, the sequences had to be regenerated using ProteinMPNN.
started from a random sequence and used a Monte Carlo sequence search coupled with structure prediction by AlphaFold to design cyclic homo-oligomers. They validated designs experimentally and showed that ProteinMPNN can rescue previously failed designs made using Rosetta or AlphaFold.
built on recent deep learning protein design approaches to develop a method called ProteinMPNN. In two papers, a range of protein design problems were addressed through deep learning methods. For the inverse problem, finding a sequence that folds to a desired structure, most approaches remain based on energy optimization. Deep learning approaches such as AlphaFold and Rosettafold have made reliable protein structure prediction broadly accessible.

